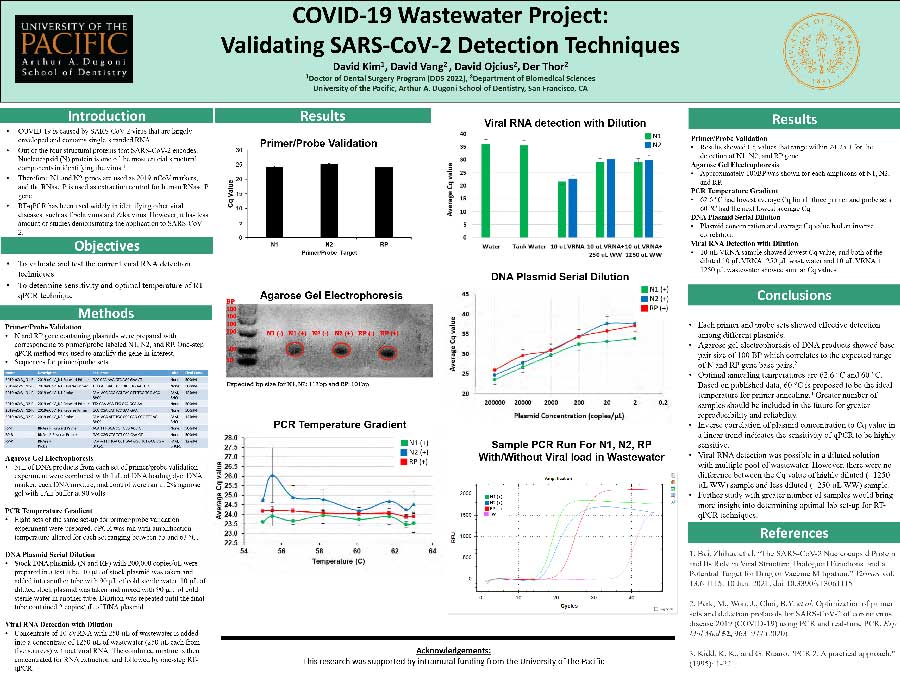
This research was done to validate and test the current viral RNA detection techniques and to determine sensitivity and optimal temperature of RT-qPCR technique.
Methods
In Primer/probe validation, N and RP gene containing plasmids were prepared with corresponding to primer/probe labeled N1, N2, and RP. qPCR method was used to amplify the gene in interest. In gel electrophoresis, 5uL of DNA products from each set of primer/probe validation experiment were ran in 2% agarose gel with TAE buffer at 90 volts. In PCR temperature gradient, qPCR was ran with amplification temperature altered for each set ranging between 55 and 63C. In Plasmid Serial Dilution, stock DNA plasmids with 200,000 copies/uL were prepared in a test tube. Dilution was repeated with 0.1x decrease until the final tube contained 2 copies/uL. In Viral RNA Detection with Dilution, concentrate of 10 cVRNA with 250 uL of wastewater is added into a concentrate of 1250 uL of wastewater (250 uL each from five sources) without viral RNA. The combined mixture is then concentrated for RNA extraction and followed by one-step RT-qPCR.
Results
Cq values ranged 24-25.1 for all genes. Approximately 100BP was shown for all amplicons. 62.6C had lowest average Cq and subsequently 60C. Plasmid concentration and average Cq value had an inverse correlation. 10 uL VRNA sample showed lowest Cq value, and both of the diluted 10 uL VRNA+250 uL wastewater and 10 uL VRNA + 1250 uL wastewater showed similar Cq values.
Conclusions
Each primer and probe sets showed effective detection among different plasmids. DNA products correlated to the expected range of base pairs of ~100BP. 62.6 and 60C are the ideal temperatures for primer annealing. Plasmid dilution demonstrated the sensitivity of qPCR to be highly sensitive. Viral RNA detection was possible in a diluted solution with multiple pool of wastewater. However, there were no difference between the Cq value of highly diluted sample and less diluted sample. Further study with greater number of samples would bring more insight into determining optimal lab set-up for RT-qPCR techniques.
Acknowledgements
This research was supported by intramural funding from the University of the Pacific
Author(s): David Kim, DDS '22
Mentor(s): NA
ADA Foundation/Colgate Dental Students Award
[embeddoc url="https://excellenceday.pacific.edu/wp-content/uploads/2023/02/David_Kim_COVID-19_Wastewater_Project_Validating_SARS-CoV-2_Detection_Techniques.pdf"]